Elijah Cole
FMs and Perturbation Prediction: Good Embeddings vs. Fancy Architectures
This post discusses results from Foundation Models Improve Perturbation Response Prediction - check out the paper for full details. All opinions are my own. Thanks to Caleb Ellington for feedback on this post.
TLDR: Fancy perturbation prediction methods (e.g. STATE, GEARS, diffusion models) don’t readily outperform FM embeddings + simple predictors (e.g. kNN).
Introduction
Perturbation modeling is a very hot topic in biology, and is closely aligned with the push towards a virtual cell. New papers come out all the time, using the latest ideas from mainstream ML in hopes of accurately simulating how cells respond to drugs or other interventions. If successful, this would be a huge deal for drug discovery and basic biology. However, it’s hard to say how much progress this stream of papers really represents. Unlike in e.g. computer vision, there’s no universally accepted shared leaderboard and most papers don’t perform ablation studies to isolate the important parts of their models. It’s unclear what works and what doesn’t.
In our paper, we found that FM embeddings combined with simple predictors can do really well on some perturbation tasks. What happens when we use the FM embeddings with the more complex models coming out of the community?
If you control for the perturbation embedding, fancy methods don’t beat kNN
Generative models have become popular in perturbation response modeling. The basic idea is to train some sort of generative model that allows you to sample from a distribution p(cell_state | perturbation, cellular_context).
Let’s consider two design choices here:
- How should we represent perturbations?
- Which generative model should we use to map perturbations to outcomes?
Most papers entangle these two choices, treating the perturbation encoding as if it’s an intrinsic part of the model and not a module that can be freely swapped out. This is quite different from fields like computer vision, which have built their success on pretrained components (e.g. the classic ImageNet-pretrained ResNet-50) that are used as building blocks in task-specific models. Good pretrained models are especially useful when training datasets are small - and most perturbation datasets include relatively few perturbations and biological contexts. Perhaps perturbation modeling should follow computer vision’s lead by cleanly separating pretrained embeddings from predictive models and analyzing the utility of each module in isolation.
This is precisely what we do in one of the experiments in the Supplemental Materials of our paper. We pick a representative collection of “advanced” perturbation prediction methods, create minimal implementations of them, and give each method the exact same perturbation embedding:
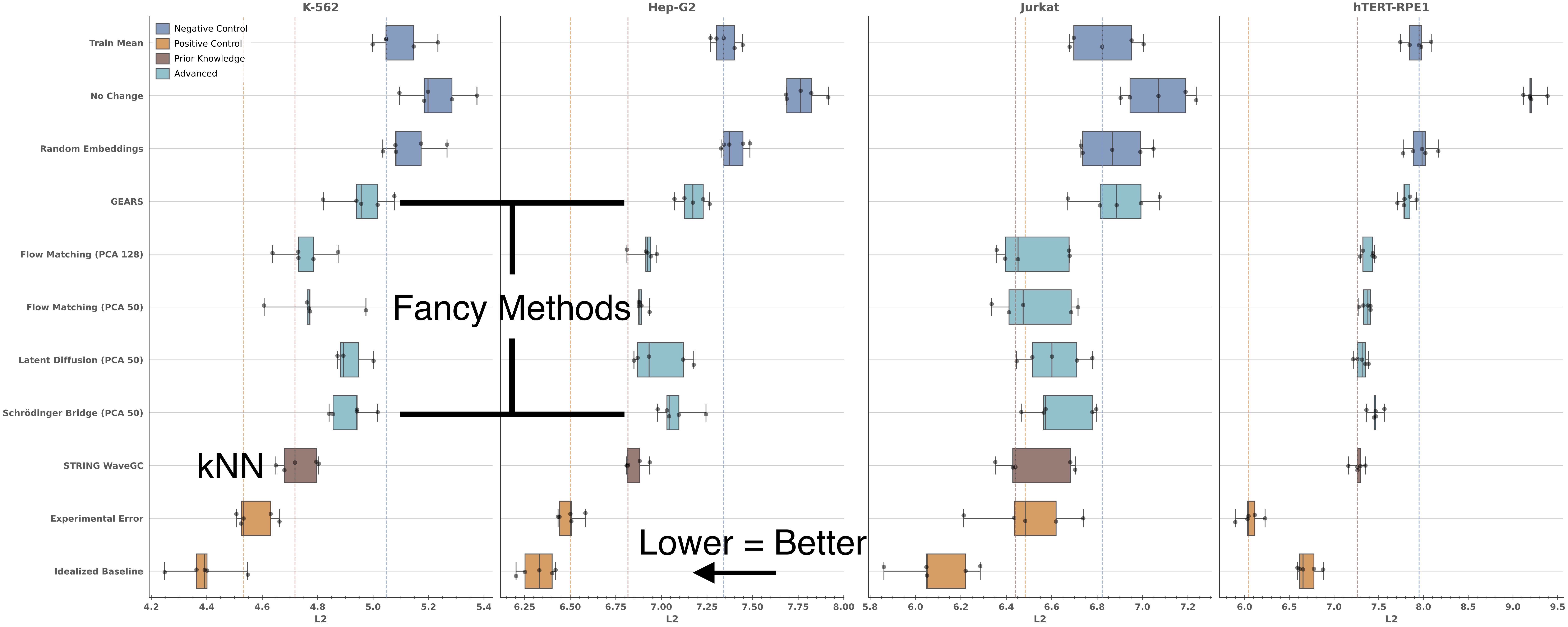
All of these models are starting with the exact same perturbation embedding, but there are clear performance differences. Flow Matching, Latent Diffusion, and Schrödinger Bridge all outperform GEARS. This is good, because it indicates that more recent architectures are providing a boost. Flow matching seems to be the best of the lot.
However, none of these methods ever convincingly outperforms kNN when it comes to predicting the average population response. This is not ideal, given how much more complex and expensive these methods are to run.
A few caveats:
- These “advanced methods” do not receive as much hyperparameter tuning as kNN. This is simply because they have more hyperparameters and are more costly to train, often requiring hours or days for a single run. It may be that there exist hyperparameters that allow these models to outperform kNN, but we did not find them. The code for this experiment is available here, please feel free to give it a try!
- We’re evaluating the average response, even though some of these methods make single-cell predictions. Perhaps the models are doing something impressive in capturing other parts of the distribution but not predicting the mean well. This is possible, but we’d need to evaluate distributional metrics with which this simple kNN model is not compatible.
The previous experiment held the embedding constant and varied the architecture in an “unseen perturbation” setting. Next, we ask a different question: how do embedding-driven baselines compare to STATE in the “unseen biological context” setting?
STATE does not consistently beat simple baselines
There has been a lot of excitement around the STATE paper from the Arc Institute, so we were excited to include it in our study. (We mention our STATE experiments briefly in the main paper, but full details and figures can be found in the Supplemental Materials.)
However, STATE considers a different perturbation prediction problem formulation than we do:
- We focus on making predictions for unseen perturbations within a given cell line. STATE makes predictions for previously seen perturbations in unseen cell lines.
- Our work focuses on the average response over cells, while STATE predicts a distribution of responses. We therefore could not include STATE in our core comparisons without modifying how STATE works.
Instead of adapting STATE to our setting, we took our embedding-centric approach to STATE’s setting. Specifically, we introduced the following simple models:
- kNN
- Inference Procedure: For each control cell
y, make a prediction by finding the 3 training cell lines with the highest cosine similarity and returning the average perturbation effect for those cell lines. - Keep in mind that this is a different kNN than we used in the previous section!
- This model actually doesn’t use a perturbation embedding, but we thought it was a sensible baseline to add to the mix.
- Inference Procedure: For each control cell
- MLP
- Inference Procedure: For each control cell
y, make a predictiony+mlp(x)wherexis an embedding of the perturbation of interest. - We try two versions of this model - one where we use an FM embedding and one where we use a one-hot encoding of the perturbation. This ablation tells us whether the FM embeddings are really doing any work. This turns out to make little difference most of the time, which is not too surprising given that all of the perturbations have been seen during training.
- Inference Procedure: For each control cell
Okay, now we are in business - the kNN and MLP models produce a population of perturbed cells, and can be directly compared to STATE. Keep in mind that MAE is “lower is better” while the other metrics are “higher is better.”
For the Tahoe dataset, STATE significantly outperforms baselines in only one of three metrics:
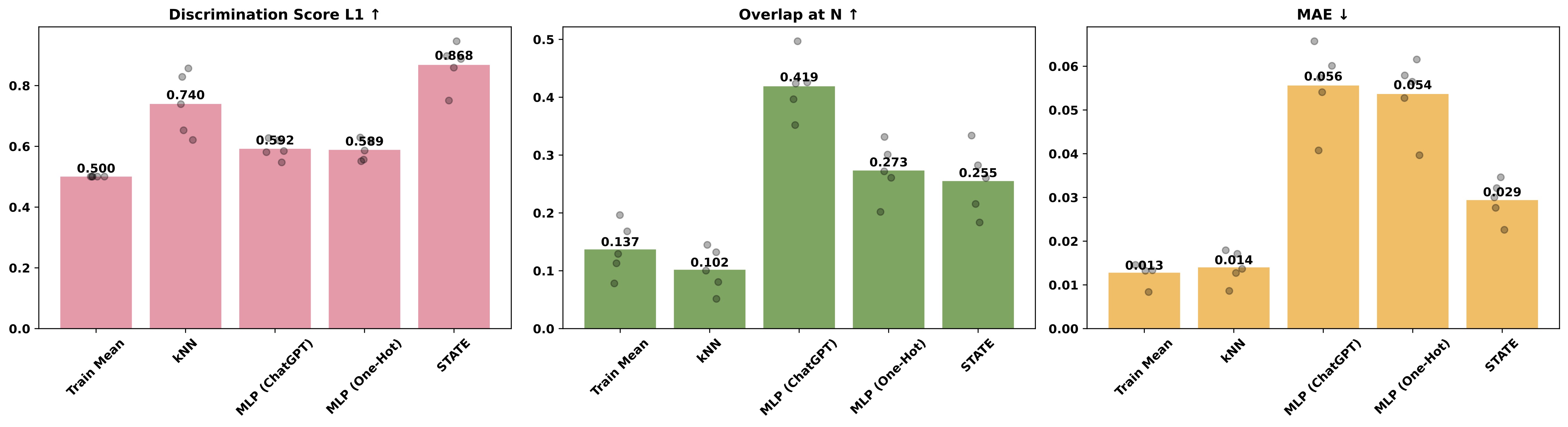
The same is true for the Essential dataset:
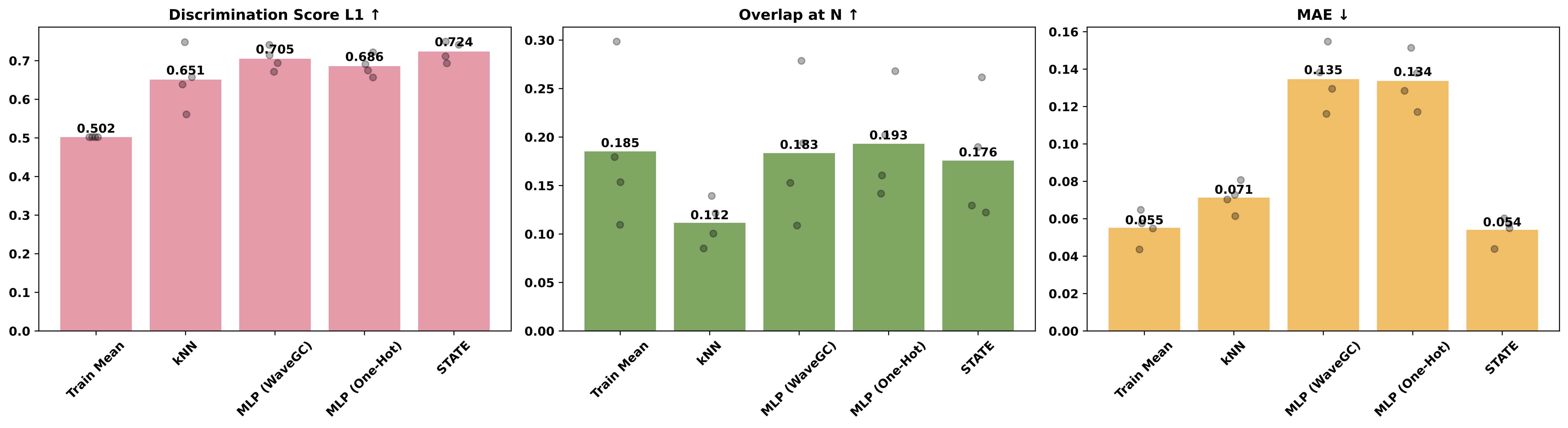
Overall, STATE (right-most bar) convincingly outperforms simple baselines in only two out of six cases. On the MAE metric, no method clearly outperforms Train Mean (left-most bar; the naive baseline that always predicts the mean of the training set). Our results do not support the idea that STATE provides a step-change in performance for perturbation prediction.
A few notes:
- The MLP and kNN baselines did not undergo any systematic hyperparameter tuning, so their performance may be understated.
- STATE’s cross-context formulation is difficult for all models we evaluate. More work on the cross-context formulation is clearly needed.
- Our numbers don’t exactly match up with the STATE paper, but they should since the settings are identical. We used the official STATE codebase for all of these experiments, so to the best of my knowledge discrepancies are due to mismatches between the official STATE codebase and the paper. All code for this analysis is available here, including the copy of the STATE codebase we used.
Concluding Thoughts
I think perturbation modeling would benefit from more focus on representation learning, and a clean separation between learned embeddings and predictive models built on top of those embeddings. Ideally we’d have an “ImageNet-pretrained ResNet-50” for perturbation modeling, a reliable pretrained model we can always reach for when building something new. We show in the paper that some embeddings perform extremely well for some datasets, but we have yet to see the emergence of globally useful perturbation representations. Overall, our results indicate that good embeddings are more important than fancy models trained on top of those embeddings. It’s important to remember that, no matter how many cells you have, perturbation modeling is very much a few-shot problem in terms of the number of perturbations and biological contexts. Training a complex model for each perturbation dataset individually would require more data than we typically have.
Key Takeaway: In a few-shot problem like perturbation modeling, embeddings do the heavy lifting for generalization, not the complexity of the predictor on top.